Key Features & Benefits
Broad model portfolio – Covers eosinophilic CRS (papain), fungal protease-induced CRS (Aspergillus), superantigen-associated CRS (SEB), and classic allergic rhinitis (OVA).
Multiple strains – C57BL/6 and BALB/c available to suit different genetic backgrounds and Th1/Th2 biases.
Comprehensive endpoints – Body weight, nasal lavage cell counts (eosinophils, total cells), serum IgE (total and OVA-specific), nasal scratching behavior, nasal mucosa histopathology (HE), cytokine profiling (IL-33, Th2 cytokines).
Translational value – Ideal for testing corticosteroids, antihistamines, biologics (anti-IgE, anti-IL-5, anti-IL-4Rα), and novel immunomodulators.
IND-ready data packages – Studies can be conducted in accordance with GLP principles.
Technical Data & Validation
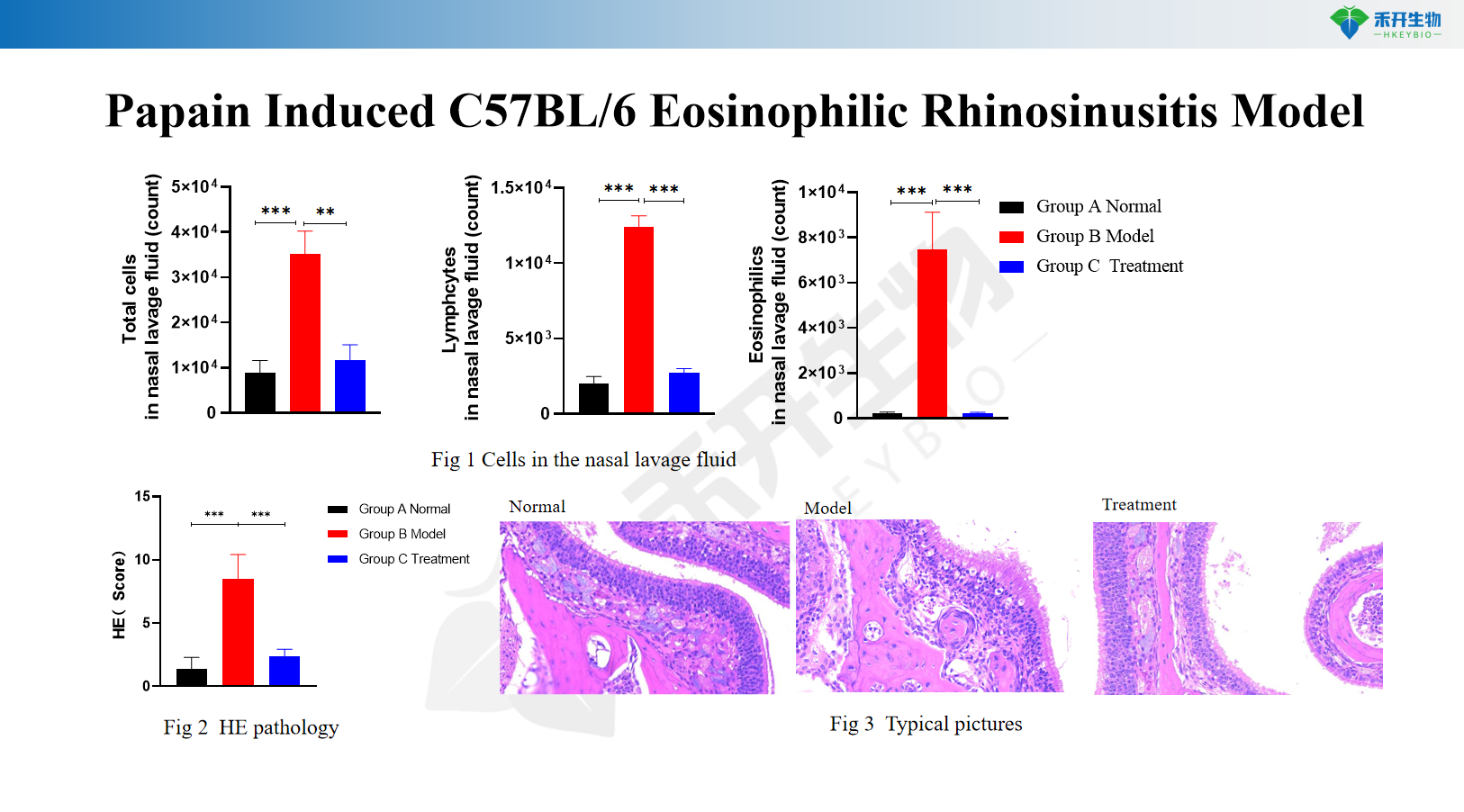
Papain Induced C57BL/6 Eosinophilic Rhinosinusitis Model
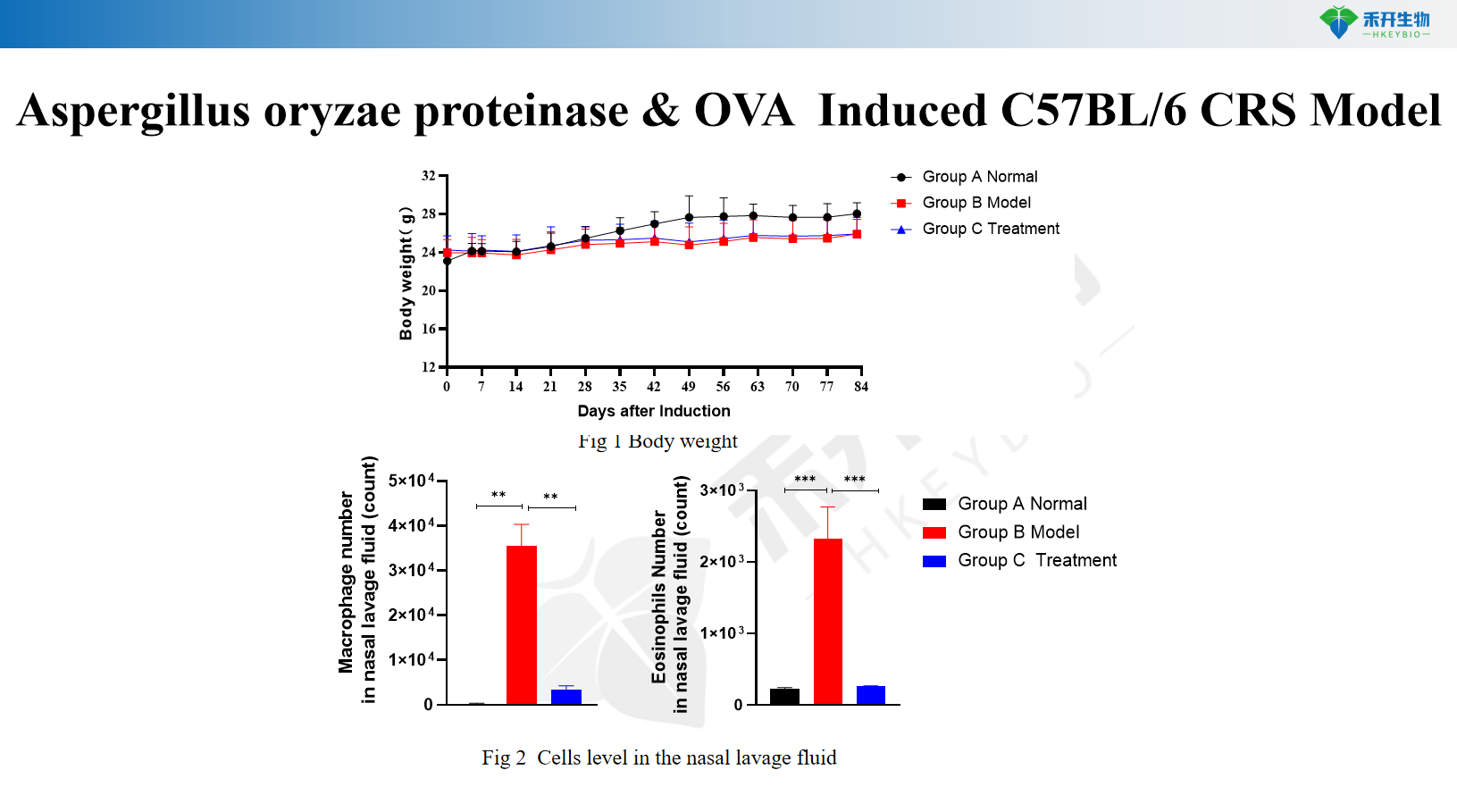
Aspergillus oryzae proteinase & OVA Induced C57BL/6 CRS Model
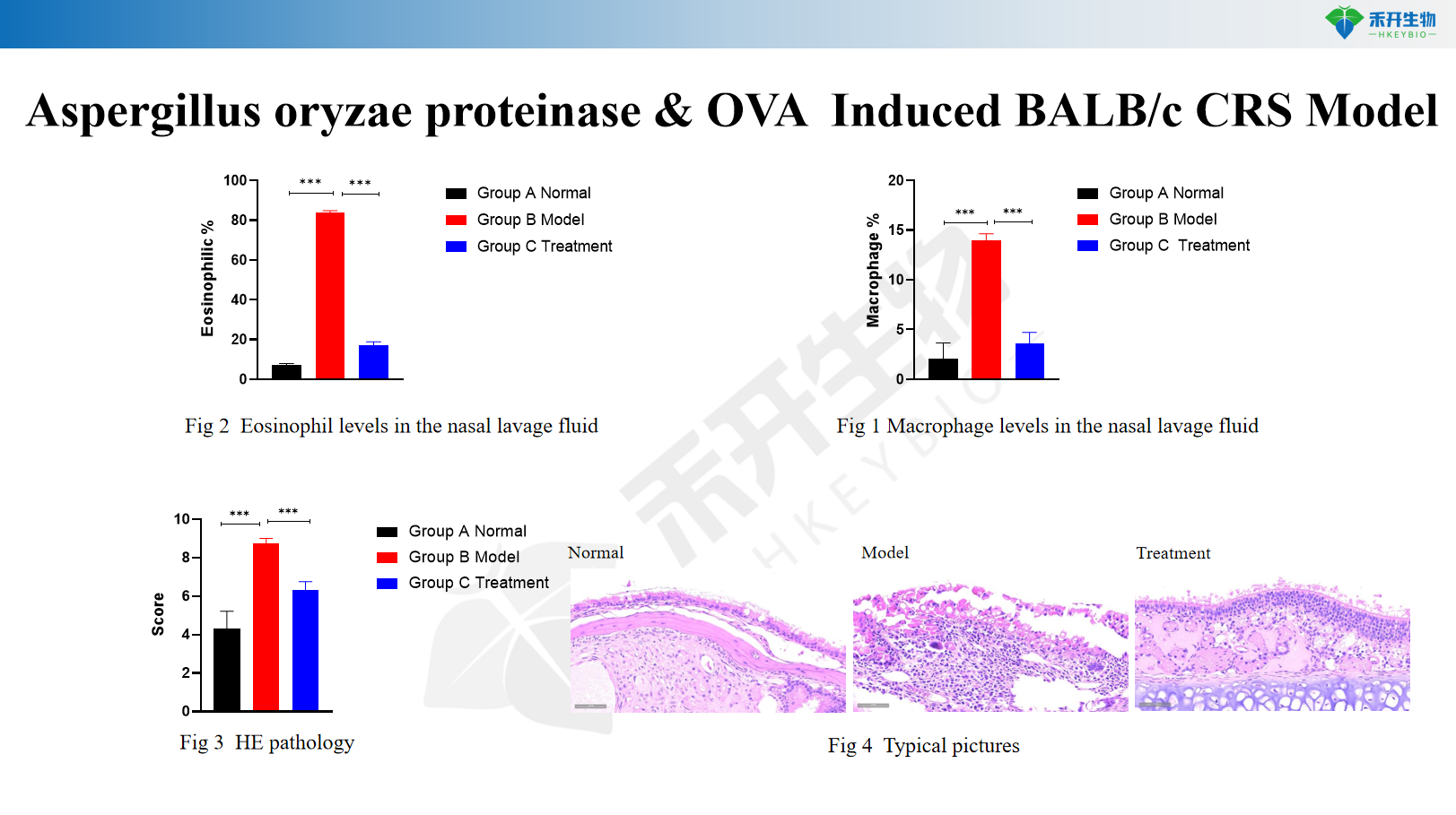
Aspergillus oryzae proteinase & OVA Induced BALB/c CRS Model
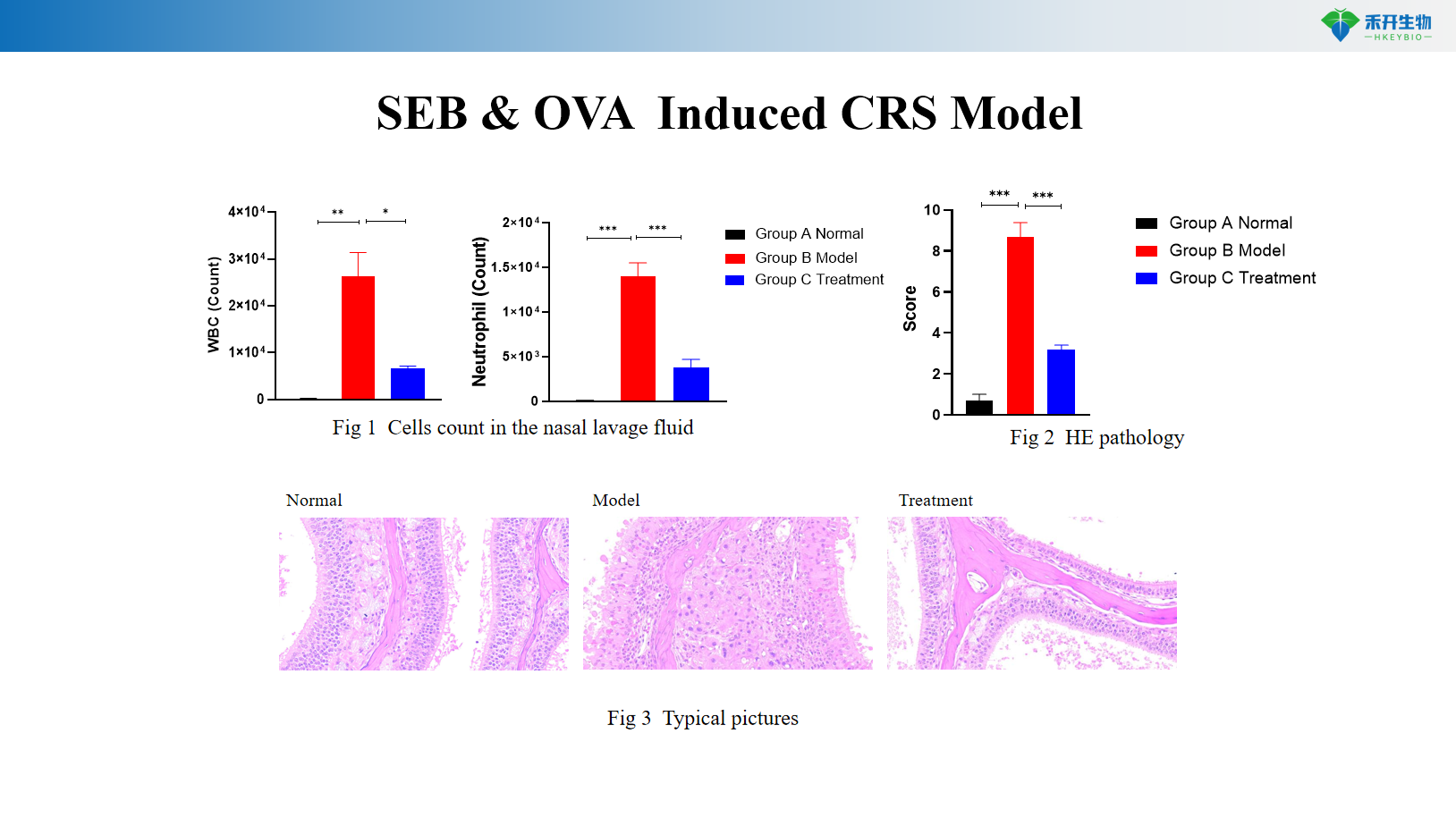
SEB & OVA Induced CRS Model
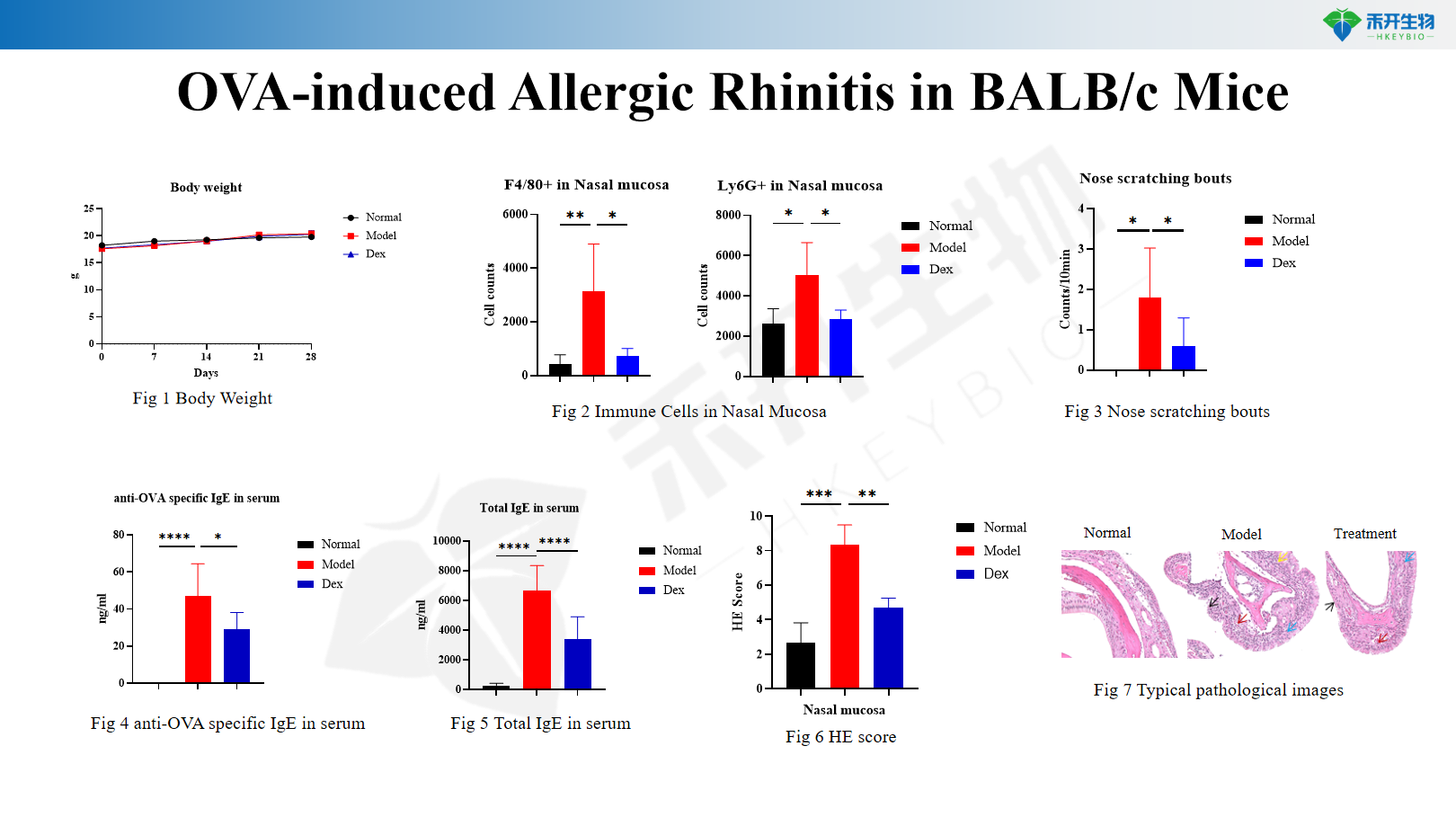
OVA-induced Allergic Rhinitis in BALB/c Mice
Applications
• Efficacy testing of intranasal and systemic corticosteroids, antihistamines, and decongestants
• Evaluation of biologics targeting Th2 pathways (anti-IL-4Rα, anti-IL-5, anti-IL-13, anti-IgE)
• Target validation for epithelial-derived cytokines (TSLP, IL-33, IL-25) and protease-activated pathways
• Biomarker discovery (IgE, eosinophil peroxidase, cytokine signatures)
• IND-enabling pharmacology and toxicology studies
Model Specifications
Parameter | Specification |
Species/Strain | Mouse (C57BL/6, BALB/c) |
Induction method | Papain (protease); Aspergillus protease + OVA; SEB + OVA; OVA + alum |
Study duration | 3–6 weeks (sensitization + challenge phases) |
Key endpoints | Body weight, nasal lavage cell counts (total and differential), serum total IgE and OVA-specific IgE, nasal scratching behavior (allergic rhinitis), nasal mucosa histopathology (HE scoring for inflammation, eosinophilic infiltration, goblet cell hyperplasia), cytokine levels (IL-4, IL-5, IL-13, IL-33) in nasal tissue/lavage |
Data package | Raw data, analysis reports, nasal lavage cytology, ELISA results, histology slides, behavioral data, bioinformatics (optional) |
❓ Frequently Asked Questions
Q: How do I choose the right model for my drug candidate?
A: For eosinophilic CRS, papain or Aspergillus protease models are recommended. For superantigen-associated CRS, the SEB+OVA model is appropriate. For classic allergic rhinitis, the OVA model is the standard choice. BALB/c mice exhibit stronger Th2 responses, while C57BL/6 allow use of transgenic lines. Our scientific team can guide model selection based on your specific target.
Q: What is the role of protease activity in CRS models?
A: Proteases (papain, Aspergillus protease) disrupt epithelial tight junctions, leading to barrier dysfunction and release of epithelial cytokines (IL-33, TSLP), which drive type 2 inflammation and eosinophilic infiltration, mimicking human CRS pathophysiology.
Q: Can these models be used for IND-enabling studies?
A: Yes. Studies can be conducted in accordance with GLP principles for regulatory submissions (FDA, EMA).
Q: Do you offer customized study protocols (e.g., different allergen doses, sensitization schedules)?
A: Absolutely. Our scientific team tailors induction protocols, treatment schedules, and endpoint analyses to your specific drug candidate.



































